Snakemake on AWS ParallelCluster using Slurm
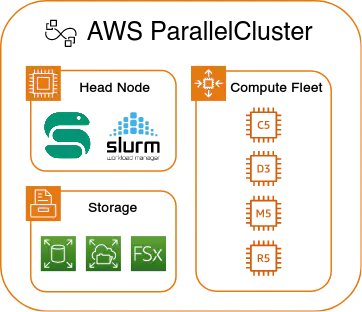
Table of Contents
tl;dr
- AWS ParallelCluster is a great way to run traditional HPC workflows in the cloud.
- AWS ParallelCluster Slurm has a few peculiarities in the default configuration.
- For running Snakemake an adapted profile is available at https://github.com/cbrueffer/snakemake-aws-parallelcluster-slurm to make things work smoothly.
Running Snakemake on AWS ParallelCluster
Snakemake is a fantastic tool for orchestrating bioinformatics workflows and comes with built-in support for a variety of public cloud providers, such as Amazon AWS. However, running a workflow this way can be a challenge, not least since hands-on tutorials or more practical information are rare. An alternative that many bioinformaticians will be familiar with are high-performance computing (HPC) clusters using a job scheduler such as Slurm, since many universities and research facilities operate such systems on their own premises.
AWS ParallelCluster
AWS ParallelCluster makes it possible and relatively easy to replicate such a setting in the cloud. ParallelCluster is essentially a wrapper around more basic AWS services such as EC2 (running virtual machines), EFS and EBS (storage), and others.
Pros and Cons
Pros
- Feasible to set up for a single user or small company / research group without huge resources.
- It scales dynamically: nodes are allocated when needed and decommissioned when unused.
- Practically infinite resources: you can allocate as many nodes and use as much storage as you want.
- It makes it easy to transfer a workflow from a traditional on-premise HPC setting into the cloud.
Cons
- In a university setting you can often get time on HPC systems for free, only requiring a short project description. Cloud-based HPC will cost you actual money.
- Traditional HPC clusters have qualified systems and network administrators that keep things running smoothly. With ParallelCluster you are the systems and network administrator, for better or worse. While many things are automated, Amazon AWS is nothing if not complex and it still requires knowledge to make things work.
Setup
The first steps of using ParallelCluster are pretty well documented.
All tasks are performed with the aws-parallelcluster Python package.
Note that there are currently two versions of the ParallelCluster software, v2 and v3.
v3 is newer and has more features, but many guides and help you currently find online are based on v2.
The most visual difference is the configuration format: v3 uses YAML while v2 uses INI files.
- Configure an AWS user with the correct permissions. A basic policy is provided in the AWS documentation, but if you need access to services such as EFS more permissions will be needed.
- Set up the AWS user’s credentials using
aws configure. - Either import an SSH public key in the EC2 settings, or generate a new key pair.
- Install the
aws-parallelclusterPython package usingpipas documented. The package is also available throughcondavia the conda-forge channel, but as of this writing it is v2 only. - Set up a cluster configuration using
pcluster configure --config my-cluster-config.yaml. This starts an interactive process that various settings of the cluster, i.e. which instance type to use for the head node, which scheduler to use (Slurm or AWS Batch), how many queues, number of nodes per queue, etc. In particular sets up the necessary virtual private cloud (VPC), network infrastructure, and security groups on AWS automatically, if you tell it to do so. - Start the cluster using
pcluster create-cluster --cluster-name test-cluster --cluster-configuration my-cluster-config.yaml. This will take a few minutes; you can check progress usingpcluster describe-cluster --cluster-name test-cluster. - Log in to the head node using
pcluster ssh --cluster-name test-cluster.
Snakemake on ParallelCluster with Slurm
Peculiarities of ParallelCluster Slurm Instances
Generally using Snakemake on a ParallelCluster Slurm cluster works the same as with any other Slurm system. That said, there are some differences and drawbacks, and they are not well documented and as such, are learned through the usual cycle of trial, error, and searching the web.
The
sbatch--memargument for specifying memory requirements for a job is not supported, and if used will send nodes into theDRAINEDstate. The company Ronin has a good troubleshooting page on this topic. If you are the sole user of your cluster the easiest workaround is probably to scale jobs using the number of CPUs (--cpus-per-task).By default, a ParallelCluster comes without working job accounting, which means
sacctdoes not work. This is particularly noteworthy since many Snakemake Slurm profiles usesacctfor job monitoring. A guide for setting up Slurm accounting on ParallelCluster is available from Amazon. If you are going to use ParallelCluster regularly it probably makes sense to set it up, since the underlying database will persist across cluster instances.
Snakemake Slurm Execution Profiles
The profile I use is freely available on Github at https://github.com/cbrueffer/snakemake-aws-parallelcluster-slurm and accomodates the ParallelCluster peculiarities. It is heavily based on John Blischak’s smk-simple-slurm repository and carries the same license.
Other profiles, such as the standard Snakemake Slurm profile do not currently work in this setting out of the box.
Other Tidbits and Lessons Learned
There is a graphical frontend to ParallelCluster! It is relatively new and not particularly well advertised in the documentation, but it looks useful and can help avoid potential configuration issues mentioned in this section. It also simplifies setting up Slurm accounting.
Shared storage
/homeis shared between the head node and the compute nodes./scratchis the local storage of each node.- Other shared storage has to be explicity added to the configuration in ParallelCluster v3 (as opposed to v2 where
/sharedis a shared EBS volume).
Head node instance type
Give some thought to which instance type to select for the head node to best balance cost and performance. Since it is the one cluster node that will run for as long as the cluster is up, the instance type should not be too expensive. On the other hand it needs to be performant enough for Snakemake, job scheduling and NFS server duties, so do not select an instance type that is too underpowered. For my applications
t3.xlargewas a good compromise.Check whether the instance types you select exist in your configured AWS region. Not all instance types are available in all regions, and
pclusterwill not warn you about this.Check your limits!
AWS services that ParallelCluster uses may have limits in place, including limits on concurrent EC2 instances and virtual CPUs. Again,
pclusterwill not warn you about this. In practice this means you can configure and start a cluster that could scale up to hundreds of compute nodes, but when you start a job only a few nodes spin up while the rest of your jobs are stuck waiting for resources and you are left wondering why.If things do not work, check the troubleshooting documentation and the logs.
On the head node:
/var/log/cfn-init.log /var/log/chef-client.log /var/log/parallelcluster/slurm_resume.log /var/log/parallelcluster/slurm_suspend.log /var/log/parallelcluster/clustermgtd /var/log/slurmctld.logOn compute nodes:
/var/log/cloud-init-output.log /var/log/parallelcluster/computemgtd /var/log/slurmd.logEphemeral storage is ephemeral!
All storage that is created during cluster creation will also be destroyed when the cluster is deleted. This includes additional EFS and EBS storage defined in the cluster config, unless you explicitly create them yourself first and then include them e.g. via their
FileSystemIdfor EFS file systems!Multi-queue clusters
The
pcluster configwizard enables creating ParallelCluster Slurm clusters with multiple queues. Keep it mind that it is only possible to specify one instance type per queue.
Resources
- Snakemake profile for AWS ParallelCluster with Slurm
- ParallelCluster documentation
- pcluster package documentation
- ParallelCluster graphical frontend
- Amazon HPC Tech Shorts Youtube channel
- Amazon HPC Tech Shorts on pcluster manager GUI
- Amazon HPC Tech Shorts on pcluster v3
Edit History
- 2022-07-19 Initial version